Bridging the Seeing-Without-Naming Gap
In agriculture, correctly naming a species from its image—what we call open-ended taxonomic naming—is the starting point for intelligent agriculture, from choosing a pesticide to enforcing quarantine. Yet no existing benchmark directly tests whether Large Multimodal Models (LMMs) can perform this task. Existing agricultural datasets each cover a single domain (8–102 classes), lack links to standard databases, and evaluate only closed-set classification—leaving untested whether models can actually name what they see.
AgriTaxon fills this gap by unifying four agricultural domains—Crops, Livestock, Pests, and Weeds—into a single benchmark with 7,432 species, each linked to authoritative FAO and EPPO databases via Wikidata. We evaluate 14 LMMs under both multiple-choice (with semantically hard negatives) and open-ended naming, and introduce an LLM-as-a-Judge scoring protocol that achieves 98% agreement with domain experts.
Leaderboard
Accuracy (%) on AgriTaxon for 14 LMMs, ranked by open-ended Acc within each block. Open-ended reports both EM (exact match) and Acc (with LLM-as-a-Judge alias validation). Bold = best per column.
| Model | Release | Hard | Multi-Choice | Open-Ended | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crop | Live. | Pest | Weed | Mean | Crop | Live. | Pest | Weed | EM | Acc | |||
| Proprietary Models | |||||||||||||
 gemini-3-pro-preview gemini-3-pro-preview |
2025.11 | 9.0 | 83.7 | 95.5 | 72.5 | 78.1 | 82.5 | 43.6 | 49.4 | 30.0 | 53.0 | 44.0 | 51.2 |
 doubao-seed-2-0-pro doubao-seed-2-0-pro |
2026.02 | 6.0 | 81.3 | 88.8 | 70.9 | 76.6 | 79.4 | 45.3 | 34.8 | 39.6 | 56.6 | 44.1 | 48.6 |
 gemini-3-flash-preview gemini-3-flash-preview |
2025.12 | 8.7 | 84.9 | 95.1 | 72.7 | 79.1 | 83.0 | 22.4 | 37.6 | 22.9 | 50.4 | 33.4 | 48.1 |
 doubao-seed-2-0-lite doubao-seed-2-0-lite |
2026.02 | 5.6 | 78.1 | 87.6 | 70.1 | 74.7 | 77.6 | 37.3 | 33.7 | 30.5 | 48.8 | 37.6 | 44.0 |
 gpt-5 gpt-5 |
2025.08 | 8.4 | 79.1 | 92.9 | 68.8 | 73.3 | 78.6 | 30.6 | 34.3 | 17.2 | 36.2 | 29.6 | 37.6 |
 gpt-5-mini gpt-5-mini |
2025.08 | 7.7 | 71.6 | 87.5 | 62.7 | 64.6 | 71.6 | 28.4 | 26.4 | 9.8 | 23.6 | 22.0 | 27.4 |
 claude-haiku-4-5 claude-haiku-4-5 |
2025.10 | 4.7 | 58.1 | 74.5 | 52.9 | 56.2 | 60.4 | 12.5 | 19.7 | 4.0 | 8.6 | 11.2 | 14.7 |
| Open-Source Models | |||||||||||||
 kimi-k2.5 kimi-k2.5 |
2026.01 | 2.1 | 75.0 | 86.0 | 65.8 | 68.0 | 73.7 | 29.8 | 31.0 | 20.0 | 39.2 | 30.0 | 38.0 |
 glm-4.6v glm-4.6v |
2025.12 | 6.2 | 66.7 | 77.0 | 57.9 | 58.6 | 65.1 | 33.2 | 20.7 | 10.4 | 26.6 | 22.7 | 30.1 |
 qwen3-vl-235b-a22b qwen3-vl-235b-a22b |
2025.09 | 2.2 | 66.4 | 81.5 | 63.2 | 62.5 | 68.4 | 23.5 | 23.9 | 13.5 | 25.8 | 21.7 | 27.5 |
 qwen3.5-397b-a17b qwen3.5-397b-a17b |
2026.02 | 9.2 | 70.5 | 85.4 | 65.8 | 65.8 | 71.9 | 25.3 | 22.8 | 13.8 | 24.8 | 21.7 | 26.8 |
 qwen3-vl-30b-a3b qwen3-vl-30b-a3b |
2025.10 | 2.2 | 59.2 | 70.1 | 55.9 | 54.6 | 59.9 | 19.9 | 24.2 | 5.3 | 14.6 | 16.0 | 22.5 |
 glm-4.6v-flashx glm-4.6v-flashx |
2025.12 | 5.8 | 59.3 | 71.3 | 55.4 | 55.1 | 60.3 | 21.6 | 20.7 | 5.7 | 12.7 | 15.2 | 19.6 |
 qwen3.5-35b-a3b qwen3.5-35b-a3b |
2026.02 | 4.2 | 67.1 | 80.9 | 61.4 | 61.7 | 67.8 | 11.3 | 17.4 | 3.9 | 12.6 | 11.3 | 17.4 |
Hard = AgriTaxon-Hard subset (≤2 models correct, 1,052 samples). EM = exact match after normalization. Acc = LLM-as-a-Judge alias validation (98% expert agreement).
📬 Submit to Leaderboard — If you would like your model or method to appear on this leaderboard, please contact us at zengx@nercita.org.cn with your evaluation results.
Key Findings
- Taxonomic confusion dominates errors (63%) — models identify the correct biological family but fail at genus- or species-level discrimination.
- 2× popularity-driven gap — popular species (>10K monthly Wikipedia views) are recognized 2.2× more accurately than obscure ones (≤100 views).
- Agentic cropping more than doubles accuracy — equipping a model with a simple image-crop tool lets it zoom into diagnostic regions, boosting accuracy on the hardest samples from 4.7% to 10.4%.
- Text-semantic distractors are hardest — taxonomically similar distractors drop accuracy from 99% (random) to 9%, confirming that label-space plausibility governs difficulty more than visual similarity.
Explore the Embedding Space
7,437 species embedded by Qwen3-Embedding-0.6B and projected via t-SNE. Scroll to zoom, drag to pan, hover for details, click to open Wikipedia.
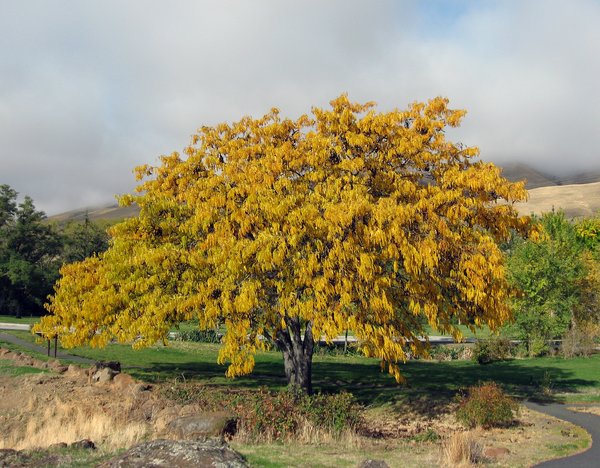
Error Type Examples
Representative examples from the human-annotated 75 Acc-error cases (Gemini 3 Pro Preview, open-ended evaluation).


How AgriTaxon Works
Authority-grounded data collection. We query Wikidata for species that carry both an authoritative database identifier (FAO Ecocrop, FAO DAD-IS, or EPPO) and a Wikimedia Commons image, forming a traceable authority chain for every label.
Cross-domain coverage. The 48,950+ EPPO entries—mixing pests, weeds, pathogens, and host plants—are classified via LLM into pest and weed tracks. All images undergo resolution filtering (≥224px) and visual content validation.
Dual evaluation protocols. Multiple-choice uses semantically hard negatives (top-3 similar species by text embedding); open-ended requires free-form species name production. An LLM-as-a-Judge protocol handles alias matching (scientific names, common names, synonyms) with 98% expert agreement.
Potential Applications
AgriTaxon is designed to support a broad range of research directions across the multimodal AI and agricultural informatics communities.
Open-Ended Visual Recognition
A testbed for models that must produce free-form species names rather than selecting from a fixed label set.
Long-Tail Understanding
Popularity metadata enables controlled study of how accuracy degrades for rare, economically important organisms.
Retrieval-Augmented Generation
Authority-grounded labels (FAO, EPPO, Wikidata QIDs) provide natural retrieval anchors for augmenting LMMs.
Agentic Reasoning
Tool use (e.g., image cropping) significantly boosts accuracy, motivating system-level augmentation research.
Agricultural AI Deployment
Pest surveillance, quarantine enforcement, crop variety verification, and livestock breed identification.
Fine-Grained Classification
Semantically hard negatives and cross-domain coverage make a challenging FGVC benchmark.
Licensing & Access
AgriTaxon is publicly available and free for academic and research use.
- Source code, prompt templates & documentation (this repository): MIT License.
- Benchmark metadata & annotations (species labels, evaluation splits, distractor sets): released under CC BY 4.0.
- Images: sourced from Wikimedia Commons under their respective Creative Commons licenses (predominantly CC BY and CC BY-SA).
- Authority identifiers: Wikidata QIDs are available under CC0; FAO and EPPO identifiers are used for reference linking only.
The dataset is hosted on Hugging Face at Xin1818/AgriTaxon and can be downloaded freely without registration.
Getting Started
# Download the dataset from Hugging Face pip install huggingface_hub huggingface-cli download Xin1818/AgriTaxon --repo-type dataset --local-dir dataset